In this vignette, you will learn everything you need to know to get started implementing numerical algorithms using {anvl}. If you have experience with JAX in Python, you should feel right at home.
The AnvlArray
We will start by introducing the main data structure, which is the
AnvlArray. It is essentially like an R array, with some
differences:
- It supports more data types, such as different precisions or unsigned integers.
- The array is managed by a specific backend (which we will ignore for now) and can live on different devices, such as CPU (aka host) or a GPU.
- 0-dimensional arrays can be used to represent scalars.
We can create an AnvlArray from R objects using
nv_array(). Below, we create a 0-dimensional array (i.e., a
scalar) that holds a 16-bit integer living on the CPU.
## AnvlArray
## 1
## [ CPUi16{} ]Note that for the creation of scalars, you can also use
nv_scalar() as a shorthand to skip specifying the shape and
omit specifying the device, as CPU is the default.
x <- nv_scalar(1L, dtype = "i16")
x## AnvlArray
## 1
## [ CPUi16{} ]We can also create higher-dimensional arrays, for example a
2x3 array with single-precision floating-point numbers.
Without specifying the data type, it will default to "f32"
for R doubles, "i32" for integers, and "bool"
for logicals.
## AnvlArray
## 1 3 5
## 2 4 6
## [ CPUi32{2,3} ]You can extract the object’s properties using getter methods.
dtype(y)## <i32>
shape(y) # or dim()## [1] 2 3
device(y)## <CpuDevice(id=0)>AnvlArrays have value semantics and are (with an
exception we cover later) never modified in-place.
y2 <- y
y2[1, 1] <- 99L
y2[1, 1]## AnvlArray
## 99
## [ CPUi32{} ]
y[1, 1]## AnvlArray
## 1
## [ CPUi32{} ]Note that such subset assignment always copies – unlike plain R,
where y[i] <- val can be done in place when
y has only one reference. This only applies to
eager execution; inside a jit()-compiled function
(covered in the Just In Time
Compilation section below) the XLA compiler can fuse the update with
surrounding operations and elide the copy entirely. See the Eager-mode
subset-assignment always copies section of the efficiency vignette
for the implications and how to work around it.
The as_array() function allows to convert
AnvlArrays back to R objects, which involves copying the
data. Note that for 0-dimensional arrays, the result is an R vector of
length 1, as R arrays cannot have 0 dimensions.
as_array(y)## [,1] [,2] [,3]
## [1,] 1 3 5
## [2,] 2 4 6AnvlArrays can also be saved to disk and loaded back via
nv_save() / nv_read(), which use the safetensors
format – a simple, cross-framework standard also used by e.g. PyTorch
and JAX:
path <- tempfile(fileext = ".safetensors")
nv_save(list(x = x, y = y), path)
loaded <- nv_read(path)
loaded$x## AnvlArray
## 1
## [ CPUi16{} ]Transforming AnvlArrays
There are two categories of functions in {anvl} that can be used to transform arrays:
- Anvl primitives, that follow the naming scheme
prim_<op>. They define the fundamental operations that can be expressed in {anvl}. These functions are rather low-level and often lack some ergonomics such as type promotion or broadcasting. Most users will not require to use them; for an overview seevignette("primitives"), and for how to add one seevignette("extending_primitive"). - The main User API (
nv_<op>functions) and the overloaded R operators that dispatch to them. They are built on top of the primitives and either add convenience or higher-level functionality.
prim_add(y, y)## AnvlArray
## 2 6 10
## 4 8 12
## [ CPUi32{2,3} ]
prim_add(y, x)## Error in `prim_add()`:
## ! `lhs` and `rhs` must have the same tensor type.
## ✖ Got tensor<2x3xi32> and tensor<i16>.
nv_add(y, x)## AnvlArray
## 2 4 6
## 3 5 7
## [ CPUi32{2,3} ]Next we define a function that computes the output of a linear model
\(y = X \beta + \alpha\), generate some
example data and call the function. We could have also used the
overloaded %*% and + operators, but chose the
underlying nv_* function for clarity.
We simulate some training data from a univariate linear model and randomly initialize some parameters that we’ll fit later.
X <- matrix(rnorm(100), ncol = 1)
beta_true <- rnorm(1)
alpha_true <- rnorm(1)
y <- X %*% beta_true + alpha_true + rnorm(100, sd = 0.5)
plot(X, y)
X <- nv_array(X, dtype = "f32")
y <- nv_array(y, dtype = "f32")
beta <- nv_array(rnorm(1), shape = c(1, 1), dtype = "f32")
alpha <- nv_scalar(rnorm(1), dtype = "f32")
y_hat0 <- linear_model_r(X[1:2, ], beta, alpha)
y_hat0## AnvlArray
## 3.6654
## 1.3981
## [ CPUf32{2,1} ]What we have done in this section is commonly referred to as eager execution. To understand what this means, we need to differentiate it from JIT compilation, which is the primary goal of {anvl} and which we will cover next.
Just In Time Compilation
JIT stands for just-in-time compilation: instead of compiling the function ahead of time, {anvl} waits until the first call (when the input shapes and dtypes are known) and only then translates the function into a single optimized executable, which is cached for subsequent calls.
To get the most out of {anvl} in terms of performance, one should
usually jit() your functions. For example, we can
jit-compile the linear_model_r function we defined earlier.
The output is a function with the same signature that produces the same
results:
linear_model <- jit(linear_model_r)
y_hat1 <- linear_model(X[1:2, ], beta, alpha)
all(y_hat0 == y_hat1)## AnvlArray
## 1
## [ CPUbool{} ]The difference from eager mode is that jit() compiles
the whole function using XLA,
which is the same compiler that underpins frameworks like TensorFlow and
JAX. The output is an executable program that runs independently of the
R interpreter. Note that under the hood, each prim_*
function is itself a jit()-compiled function.
One central assumption about programs that are
jit()-compiled is that the R function is a pure
function, so do not rely on side effects such as manipulation of global
state within such a function. See the Tracing Contract section of the
JIT deep dive for a more thorough explanation.
At the jit boundary, plain R values are transparently converted to
AnvlArrays, so you don’t need to wrap every input in
nv_array() yourself. This applies to both nv_*
calls and your own jit()-compiled functions:
## AnvlArray
## 3
## 4
## [ CPUf32?{2} ]Note the ? after the dtype in the printed output
(f32?): it marks the dtype as ambiguous. A dtype
is ambiguous when it was inferred from an auto-converted R value rather
than pinned by the user (e.g. via
nv_array(..., dtype = "f32")), so the type-promotion rules
are allowed to coerce it more freely when combining it with a
non-ambiguous operand. See vignette("type-promotion") for
the full rules.
Note that we only auto-convert
double/integer/logicals that
are
- vectors of length 11
- arbitrary arrays or matrices
Besides AnvlArrays, jit-compiled functions can also take
plain R values as arguments without converting them to
AnvlArrays internally. Such arguments must be marked as
static. Non-static (arrayish) inputs trigger recompilation
only when the input type combination changes; static inputs trigger
recompilation for every new value.
To illustrate this, we create a jitted mean-squared error function
whose reduction is configurable – reduction = "mean"
returns a scalar loss, "sum" returns the un-normalized
total:
mse <- jit(function(y_hat, y, reduction) {
se <- (y_hat - y)^2.0
if (reduction == "mean") {
mean(se)
} else {
sum(se)
}
}, static = "reduction")
mse(linear_model(X, beta, alpha), y, reduction = "mean")## AnvlArray
## 1.3679
## [ CPUf32{} ]
mse(linear_model(X, beta, alpha), y, reduction = "sum")## AnvlArray
## 136.7889
## [ CPUf32{} ]jit-compiled functions can also be called inside other
jit() calls. When this happens, the inner function is not
compiled and executed separately – instead, they are compiled together.
We combine linear_model and mse into a jitted
model_loss:
model_loss <- jit(function(X, beta, alpha, y) {
y_hat <- linear_model(X, beta, alpha)
mse(y_hat, y, reduction = "mean")
})
model_loss(X, beta, alpha, y)## AnvlArray
## 1.3679
## [ CPUf32{} ]To get a better understanding of how jit() works see
vignette("jit"). For a detailed discussion of when to
prefer eager vs. jit mode, see vignette("efficiency").
Automatic Differentiation (AD)
Another central feature of {anvl} is its ability to differentiate functions. Currently, we only support reverse-mode AD and no higher-order derivatives, but this will hopefully be added in the future. To showcase the automatic differentiation capabilities, we will use gradient descent to fit the linear model to the training data we simulated earlier – although one would usually do this by solving the normal equations of course.
Using the gradient() transformation, we can
automatically obtain the gradient function of model_loss
with respect to a subset of its arguments that we specify via
wrt. These must be AnvlArray inputs and not
static values. The resulting model_loss_grad has the same
signature as model_loss, but returns a named list of
gradients – one entry per argument listed in wrt:
model_loss_grad <- jit(gradient(
model_loss,
wrt = c("beta", "alpha")
))
model_loss_grad(X, beta, alpha, y)## $beta
## AnvlArray
## -0.0792
## [ CPUf32{1,1} ]
##
## $alpha
## AnvlArray
## 2.1543
## [ CPUf32{} ]Finally, we define the update step for the weights using gradient
descent. We group the parameters into a weights list that
the function both accepts and returns – this shows that inputs and
outputs of a jit()-compiled function can be (nested) lists
of AnvlArrays, not just bare arrays:
update_weights <- jit(function(X, weights, y, lr) {
grads <- model_loss_grad(X, weights$beta, weights$alpha, y)
list(
beta = weights$beta - lr * grads$beta,
alpha = weights$alpha - lr * grads$alpha
)
})This already allows us to fit the linear model.
weights <- list(beta = beta, alpha = alpha)
lr <- 0.1
for (i in 1:100) {
weights <- update_weights(X, weights, y, lr)
}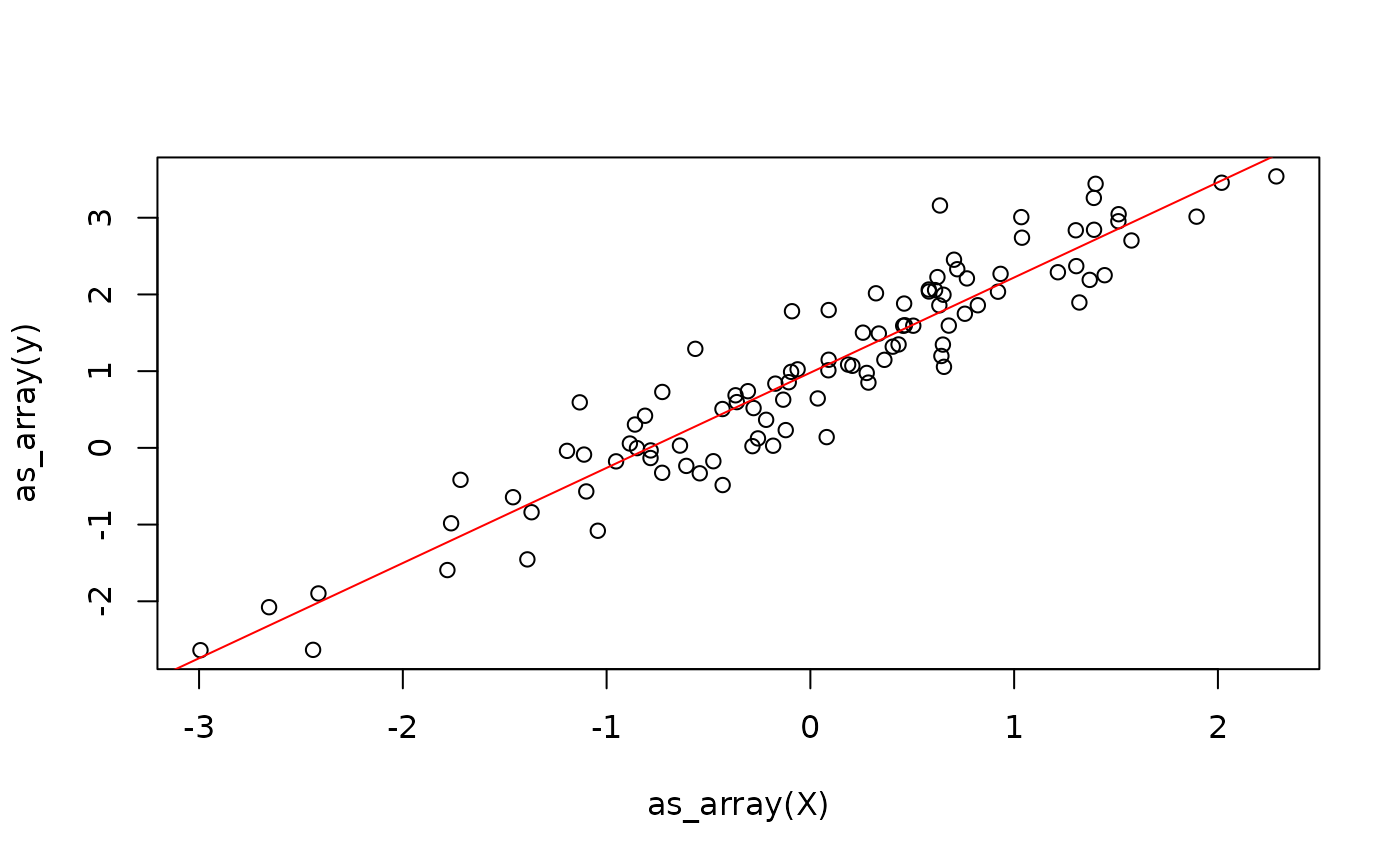
One problem with the above approach is that we are creating new
weight arrays in each iteration and throw away the previous weights,
just like we saw earlier when demonstrating subset assignment. We can
work around this using the donate argument of
jit(), which allows XLA to overwrite the inputs. See the Donation section of the efficiency
vignette for details.
Next, we will discuss control flow.
Control Flow
In principle, there are three ways to implement control flow in {anvl}:
- Embed jit-compiled functions inside R control-flow constructs, which we have seen above.
- Embed R control flow inside a jit-compiled function (we have also
seen this earlier when
msebranched onreduction). Rfor/whileloops are unrolled at trace time and Rif-statements only retain the taken branch – see the R loops are unrolled and Rifstatements pick one branch sections of the JIT deep dive. - Use special control-flow primitives provided by {anvl}, such as
nv_while()andnv_if().
Which solution is best depends on the specific use case. The first
two have already been demonstrated, so we focus on
nv_while() here. It is not like a standard while loop,
because {anvl} is purely functional. The function takes in:
- An initial state, which is a (nested) list of
AnvlArrays. - A
condfunction, which takes as input the current state and returns a logical flag indicating whether to continue the loop. - A
bodyfunction, which takes as input the current state and returns a new state.
train_while <- jit(function(X, beta, alpha, y, n_steps, lr) {
nv_while(
list(beta = beta, alpha = alpha, i = 0),
\(beta, alpha, i) i < n_steps,
\(beta, alpha, i) {
grads <- model_loss_grad(X, beta, alpha, y)
list(
beta = beta - lr * grads$beta,
alpha = alpha - lr * grads$alpha,
i = i + 1L
)
}
)
})
train_while(X, beta, alpha, y, nv_scalar(100L), lr = 0.1)## $beta
## AnvlArray
## 1.2408
## [ CPUf32{1,1} ]
##
## $alpha
## AnvlArray
## 0.9801
## [ CPUf32{} ]
##
## $i
## AnvlArray
## 100
## [ CPUf32?{} ]The same approach works analogously for if-statements,
where the {anvl} primitive nv_if() is available.